RESULTS:
In the first experimental run, 39 lines were tested for the relation between fitness (as reflected in the number of offspring produced by the females in a set) and recombination (as reflected in the proportion of recombinant progeny). As described earlier in the methods section, three sets of three females were taken from each of the 39 lines, to a total of 117 sets (see figure 8 in methods section). Of these, 116 were scored while 1 was lost. The data thus obtained was visualized in two ways, first by graphing the fitness (number of flies) of each vial against the total amount of recombination r (proportion of recombinants) in that vial, and then by graphing the mean recombination R in each line against the total fitness (sum of flies in all the three vials per line) of each line. The two graphs are as follows:
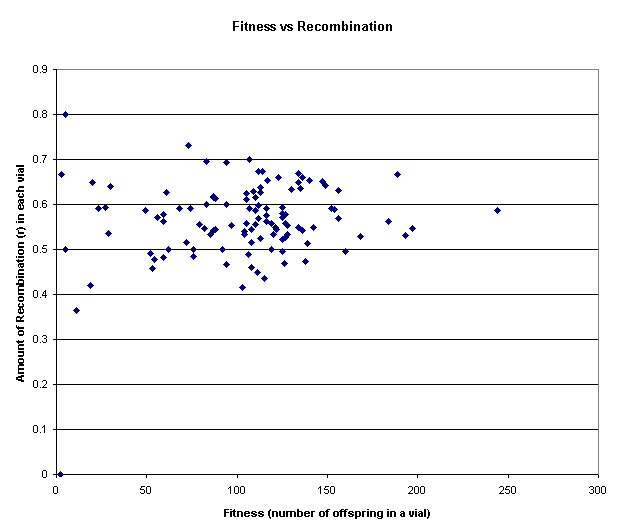
Figure 9: The above graph depicts the relation between fitness and recombination on a ‘vial by vial’ basis. On x-axis is the fitness, reflected in the total number of offspring in that vial, while on y-axis is the amount of recombination r, reflected in the number of recombinant progeny in each vial. As is quite apparent from the distribution of data points in the two dimensional plane, the slope of the data points does not point downwards as was theorized, nor is the slope as steep as might be expected in case of a true correlation between fitness and recombination.
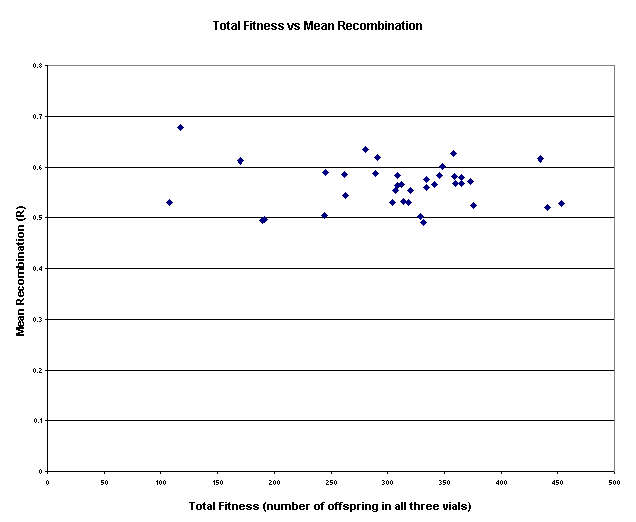
Figure
10: The
above graph depicts the relation between the total fitness and mean
recombination for the 37 lines of flies bearing the mutagenized CyO chromosome on a
wildtype background. On x-axis is the total fitness for each line as reflected
in the total number of flies in all the three vials for that line, while on
y-axis is the mean amount of recombination R for that line (the
average of three recombination scores). As is apparent from the distribution of
data points, the slope of the data points is pointing downwards, however it is
very slight.
Besides the above graphs, the correlation between fitness and recombination was also calculated, that too in two ways: first on a ‘vial by vial basis’, and then for the ‘line means’. When weighted by the inverse of sampling variance for r, the effective number of vials turned out to be 99. The value of correlation was found to be 0.084. To test whether or not this correlation was due to pure chance, a null distribution of correlation values was generated through randomizations, and 39% (P<0.4) of the times, the randomly generated correlation values were as large or larger than the correlation we obtained.
Likewise, the effective number of genetic lines, when weighted by the inverse of sampling variance for R, turned out to be 34. The value of correlation was found to be –0.006. As before, when a null distribution of correlation values was generated through randomizations, it turned out that 97% (P<0.97) of the randomly generated correlation values were as large or larger than the value we obtained.
Thus both analyses yielded far higher probabilities of our data being an artifact of chance, than the cutoff value of 5% (P<0.05), required for statistical significance.