
Mario Ventresca
Postdoctoral Fellow, University of Toronto
I am currently involved in three very large projects, briefly described below, that involve various exceptional international collaborators:
Pandemic Disease Mitigation
 The World Health Organization defines a pandemic as the worldwide spread of a new disease. Devastating pandemics such as
the Spanish Flu (H1N1), smallpox, cholera and the Black Death have occurred throughout history and it is just a
matter of time before another pandemic arises. Predicting the exact disease, its method(s) of transmission or when
exactly it will happen, is extremely unlikely. However, investigating effective contingency plans to mitigate the pandemic is possible through the use of mathematical or simulation-based models.
In this work, we aim to construct intelligent software for use by public health officials that is capable of automatically
proposing a portfolio of epidemiologically viable, economically feasible and logistically sound mitigation strategies given
a variety of realistic constraints.
The World Health Organization defines a pandemic as the worldwide spread of a new disease. Devastating pandemics such as
the Spanish Flu (H1N1), smallpox, cholera and the Black Death have occurred throughout history and it is just a
matter of time before another pandemic arises. Predicting the exact disease, its method(s) of transmission or when
exactly it will happen, is extremely unlikely. However, investigating effective contingency plans to mitigate the pandemic is possible through the use of mathematical or simulation-based models.
In this work, we aim to construct intelligent software for use by public health officials that is capable of automatically
proposing a portfolio of epidemiologically viable, economically feasible and logistically sound mitigation strategies given
a variety of realistic constraints.
The project can be decomposed into three main phases. (1) Construction of the population: given publicly available census
information, a representative heterogeneous contact network of the individuals in the geographic area under consideration is created.
(2) Identification of a critical set: from the perspective of the contact network, we aim to minimize the expected potential
number of individuals whom can become infected by effectively disconnecting the network, and provide provably accurate approximation algorithms to do so.
(3) Creation of policy portfolio: using machine learning and data mining technologies, we examine the critical set of individuals in
conjunction with census information to generate realistic mitigation scenarios (mixture of targeted vaccination, isolation, quarantine, etc) given various socio-economic and health constraints
on the system.
Reverse Engineering Complex Networks
 A complex network is a graph whose structure has some real world meaning. For instance, signal transduction in cells is
performed by a protein-protein interaction network, which represents the series of reactions that occur when two or more
proteins are combined together. It is evident then, that a comprehensive understanding of the process(es) governing the manner in
which the network's structure evolves over time can provide enormous information about the system being considered. Indeed, the potential applications
range from drug discovery to advertising. This project aims to develop a robust technology that takes as input an existing complex
network (with unknown graph model) and is able to automatically infer an algorithm that is capable of generating statistically similar structures to
the input network.
A complex network is a graph whose structure has some real world meaning. For instance, signal transduction in cells is
performed by a protein-protein interaction network, which represents the series of reactions that occur when two or more
proteins are combined together. It is evident then, that a comprehensive understanding of the process(es) governing the manner in
which the network's structure evolves over time can provide enormous information about the system being considered. Indeed, the potential applications
range from drug discovery to advertising. This project aims to develop a robust technology that takes as input an existing complex
network (with unknown graph model) and is able to automatically infer an algorithm that is capable of generating statistically similar structures to
the input network.
The past sixty years have seen complex network growth algorithms be discovered by tedious trial and error. A few standard models have been devised
that capture common scale-free and small-world properties found in many real-world networks. However, these algorithms tend to produce loosely similar
networks to the natural phenomena under consideration and thus only provide limited information to the investogator. Our approach employs
graph theory, statistics and evolutionary search algorithms to achieve results that are typically at least as good, but often far superior, to the
traditional algorithms. For instance a recent study discovered a more accurate complex network model of the cat cortex than
any previously existing in literature. A number of open theoretical questions and practical challenges remain, making this a fascinating
research direction.
Human Immunity and Vaccination
 The human immune system is highly complex and adaptive - and it needs to be in order to protect against infectious diseases such as
influenza, SARS, West Nile Virus, and so on. Yearly estimates on health and economic costs due to infectious diseases are absolutely staggering. Prophylactic
vaccination, if available, is one method that offers potential protection against such pathogens. However, infectious diseases are not
genetically stagnant and some have very high mutation rates. This allows them to evade immunity and thereby causes individuals to
become susceptible to new strains of the virus. This research aims to understand and quantify the interplay between the immune system, vaccines and pathogen
evolution.
The human immune system is highly complex and adaptive - and it needs to be in order to protect against infectious diseases such as
influenza, SARS, West Nile Virus, and so on. Yearly estimates on health and economic costs due to infectious diseases are absolutely staggering. Prophylactic
vaccination, if available, is one method that offers potential protection against such pathogens. However, infectious diseases are not
genetically stagnant and some have very high mutation rates. This allows them to evade immunity and thereby causes individuals to
become susceptible to new strains of the virus. This research aims to understand and quantify the interplay between the immune system, vaccines and pathogen
evolution.
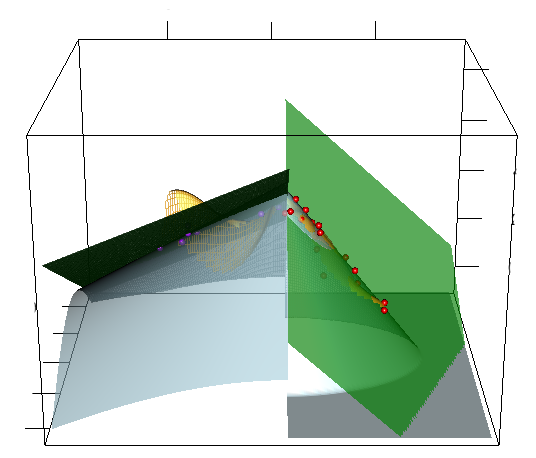
Antigenic cartography (AC) is a multi-dimensional scaling-based approach to visualizing the antigenic relationship between virus strains. Typically,
laboratory scientists create large tables of titrations of sera to the disease being considered, and AC can be used to visualize the relative differences between the
strains. For some diseases a consequence of the visualization is that the evolutionary trajectory of the virus becomes evident. For influenza (H3N2), the
visualization is a zig-zagging 2D map. Much of the research in this direction is aimed at augmenting the 2D map by including another dimension that
represents the strength of immunity of the subset of individuals being examined, thereby shedding light on the virus from the point of view of the immune system. Various tools
that provide a quantitative understanding at both the individual and population levels are in development. The long-term goal is to not only to
accurately describe pathogen evolution, but to allow for its prediction, which can have significant impact on vaccine development.