PSEUDOGENES
What are they, where do they come from and what can we learn from them?
![]()
PSEUDOGENESWhat are they, where do they come from and what can we learn from them?
|
|
Creation vs. Evolution:
|
Creation vs. Evolution:
Evolution vs. Creationism: The Debate Continues (?) It is well known fact that evolutionary scientists and creationist differ over the origins of life. Many arguments have been used by both sides to validate their views; so it is of no surprise that in the age of molecular biology that the two sides have found new ammunition -- namely pseudogenes -- to support their respective causes.
Creationist View:
The
creationist view is that sequence similarities in related species reflects the
creator’s choice to design similar species to function similarly, even at the
amino acid level. The
persistence of useless pseudogenes is a problem for evolution since the
manufacture of DNA is energetically costly to the cell.
Natural selection should therefore remove any DNA were it actually
useless. Also, the lack of
selective pressure on this neutral DNA, one would also expect that primitive
pseudogenes should be scrambled beyond recognition as a result of random
mutations (Woodmorappe, 2000). It
is not possible to state with certainty that a gene is unequivocally either a
pseudogene or a gene. Other
analysis (that have yet to be performed) in the appropriate temporal-spacial
conditions may be able to detect expression. From an evolutionary point of view, pseudogenes should be hierarchically shared between primates. However, there are examples of phylogenetic discordancy such as the immunoglobun Epsilon-pseudogene found in gorilla and man, but not chimpanzee (Ueda et al., 1985). Figure 1 illustrates that (C), which occurs in humans and orangs does not show up chimps and gorillas, the two intermediate evolutionary derivation.  Fig1: A schematic phylogeny illustrating the hierarchical (vowel) and non-hierarchical (consonant) deployment of 'shared mistakes' among five primates. Permission granted by Answers in Genesis Ltd. Brisbane, Australia. Evolutionists believe that during the course of evolution, that elements are inserted into the genome and that such episodes create a unique marker for phylogenetic analyses. Therefore pseudogene-derived phylogenies should agree well with the evolutionary trees derived from anatomic similarities. As shown below, using different pseudogenes to sort out phylogenic relationships produces differing results (Collard et al., 2000).
Table 1:
The ambiguous sharing
of “shared mistakes” shows inconsistency in a phylogenetically
derived
trees. Permission granted by Answers in Genesis Ltd. Brisbane, Australia. Evolutionists
believe that orthologous inserted-elements can
be unambigiously identified, but creationists believe that the situation isn’t
as clear cut. Orthologs
are usually far from identical as there are features which reduce the
distinctiveness of each inserted element from one another.
Alu units do not even have to be of equal
length to be judged orthologous and the
differences in the length of the poly-A tail
are
excused on the basis of vulnerability of homopolymeric
sequences to partial deletion after insertion.
Also, creationists point out that there is always some uncertainty in
aligning sequences and that there should be doubt as to whether orthologous
retropseudogenes are really located in
exactly the same position in two or more genomes.
Woodmorappe points to the fact that there are even published instances (Smit,
1996) of orthologous pairings of LINE
elements being accepted by several teams of investigators and then, upon
re-investigation, relocated hundreds of bases apart. Phylogenetically shared pseudogenes have been
termed ‘shared mistakes’ by evolutionists who conclude that such events have
an extreme improbability of occurring randomly and can only have arisen through
evolution. Creationists believe
that similar writing errors can arise independently and point to the following
evidence:
Specific target-site selection by SINE
retroelements is common as described by Miyamoto (Miyamoto, 1999).
A model which assumes constant probability of Alu
insertion cannot account for the lengthy Alu-barren intervals of host DNA (Moyzis
et al., 1989).
Alu
units often occur in clusters, to the point of aggregating at almost the same orthologous
position in different animals (Lee et al., 1998; Bailey et al., 1993). 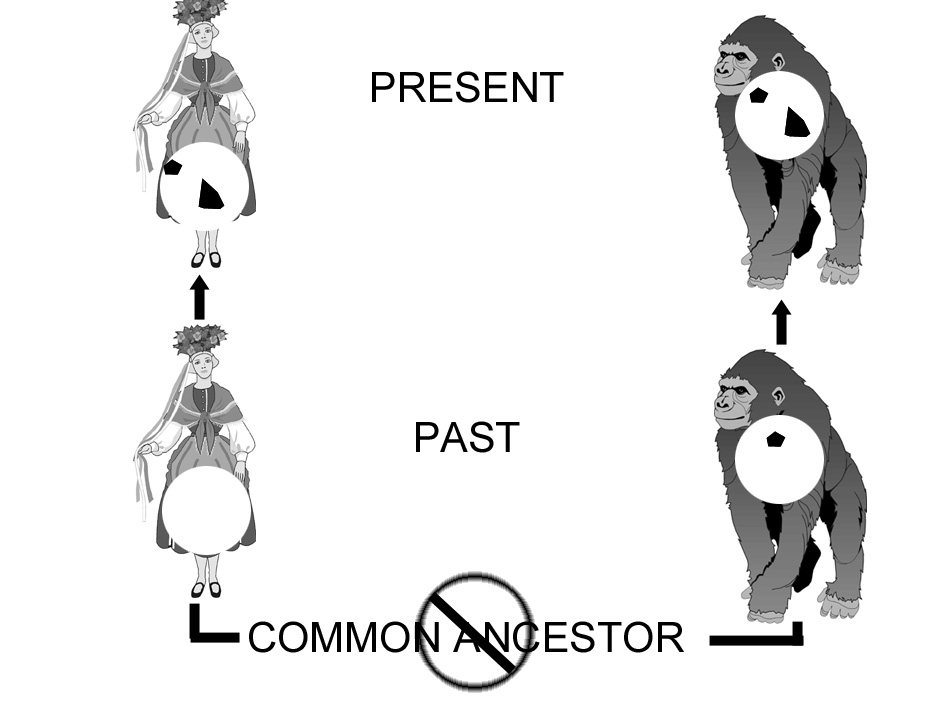 Permission granted by Answers in Genesis Ltd. Brisbane, Australia.
|
|
All Rights reserved, Copyright ©2002
T0701C - JLM349S
|